3 Chapter 3. RNA and Transcription
Suhaib Ahmad and Abdul Latif Khan
CHAPTER OUTLINE
3.3 RNA Processing in Eukaryotes
3.4 Types of RNA involved in Expression
3.5 RNA Analysis Techniques
Learning Objectives
By the end of this chapter, you will be able to:
- Explain the central dogma of molecular biology and describe the role of RNA in gene expression.
- Describe the structure of RNA and differentiate among the various types of RNA.
- Outline the key steps involved in the transcription process.
- Distinguish the processes of RNA modification and processing in eukaryotic cells.
- Identify and describe common RNA analysis techniques used in molecular biology.
The Central Dogma and Gene Expression
RNA expression is one of the basic biochemical processes that govern the vast majority of cellular activities and is considered to be the cornerstone of life. It is the process of gene transcription where DNA is copied into RNA which acts as a bridge between the genetic information in the DNA and the synthesis of proteins. This process is crucial since it helps in the management of the cellular activities, tissues and organisms as it determines the translation of genes at certain conditions and the synthesis of the molecules needed for life (Alberts et al., 2015). RNA molecules with a focus on the messenger RNAs (mRNAs) which act as a template for protein synthesis. The process of transcription, that is the conversion of DNA into RNA is a well-controlled process that makes sure that genes are expressed in the right manner at any given time and in response to various stimuli that are acting on the cell. This regulation occurs at multiple levels such as at the level of transcription and post transcription and that can affect RNA stability, localization and translation efficiency (Greco et al., 2021;Bollu et al., 2022;Chen et al., 2023).
Ribonucleic acid (RNA) is the second major type of nucleic acid found in all living cells. Like DNA, RNA is a polymer of nucleotides. Each of the nucleotides in RNA is made up of a nitrogenous base, a five-carbon sugar, and a phosphate group. In the case of RNA, the five-carbon sugar is ribose, not deoxyribose. Ribose has a hydroxyl group at the 2′ carbon, unlike RNA nucleotides contain the nitrogenous bases adenine, cytosine, and guanine. However, they do not contain thymine, which is instead replaced by uracil, symbolized by a “U.” RNA exists as a single-stranded molecule rather than a double-stranded helix.
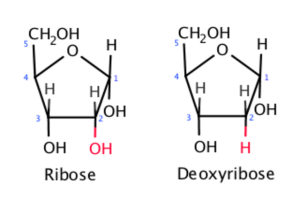
Figure 3.1: Ribose in RNA has a hydroxyl group (-OH) at the 2′ carbon, while deoxyribose in DNA lacks it. Wikimedia
RNA molecules possess unique structural features that contribute to their diverse functions:
Primary Structure
The order of nucleotides (adenine, uracil, cytosine, and guanine) in a polynucleotide. It has ribose sugar not deoxyribose which is found in DNA. The features that determine the chemical versatility of RNA include the presence of uracil in place of thymine and the 2’ hydroxyl group in ribose.
Secondary Structure
Intramolecular base pairing forms hairpins, loops, bulges and pseudo-knots. Some of the examples include the stem-loop structure that is found in tRNA and miRNA precursors. Both Watson-Crick (A-U, G-C) and non-canonical base pairings are involved in the construction of stable and functional RNA molecules.
Tertiary Structure
The three-dimensional folding of a polymer that is maintained by hydrogen bonds, van der Waals’ forces and ionic bonds. Essential for the catalytic role of ribozymes and the operation of ribosomal RNA (rRNA). Some of the tertiary interactions which are crucial for maintaining structure are coaxial stacking and RNA motifs for example kissing loops and A-minor interactions.
Quaternary Structure
RNA can interact with proteins or other RNA molecules to form larger complexes which may include the binding of RNA to RNA. Some of the examples include the ribosome, spliceosome and RNA protein granules. These structures allow for the regulation of processes such as translation and RNA splicing. In viral systems, RNA can have a structure that is associated with capsid proteins which is an example of quaternary structure of RNA with proteins. The RNA quaternary structure also has a function in the organization of phase separation and formation of RNA-protein condensates in the cells, which is very important in the organization of cellular processes.
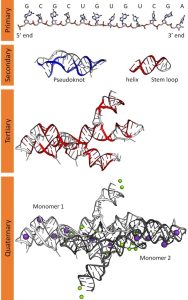
Figure 3.2: primary, secondary, tertiary, and quaternary structures of RNA. Wikimedia
In both prokaryotes and eukaryotes, the second function of DNA (the first was replication) is to provide the information needed to construct the proteins necessary so that the cell can perform all of its functions. To do this, the DNA is “read” or transcribed into an mRNA molecule. The mRNA then provides the code to form a protein by a process called translation. Through the processes of transcription and translation, a protein is built with a specific sequence of amino acids that was originally encoded in the DNA. This module discusses the details of transcription.
The Central Dogma: DNA Encodes RNA; RNA Encodes Protein
The flow of genetic information in cells from DNA to mRNA to protein is described by the central dogma (Figure 3), which states that genes specify the sequences of mRNAs, which in turn specify the sequences of proteins.
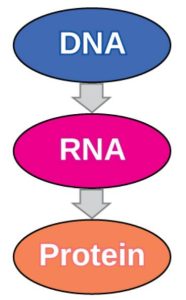
Figure 3.3: The central dogma states that DNA encodes RNA, which in turn encodes protein. OpenStax Concepts of Biology
Transcription: from DNA to mRNA
Both prokaryotes and eukaryotes perform fundamentally the same process of transcription, with the important difference of the membrane-bound nucleus in eukaryotes. With the genes bound in the nucleus, transcription occurs in the nucleus of the cell and the mRNA transcript must be transported to the cytoplasm. The prokaryotes, which include bacteria and archaea, lack membrane-bound nuclei and other organelles, and transcription occurs in the cytoplasm of the cell. In both prokaryotes and eukaryotes, transcription occurs in three main stages: initiation, elongation, and termination.
Initiation
Transcription requires the DNA double helix to partially unwind in the region of mRNA synthesis. The region of unwinding is called a transcription bubble. The DNA sequence onto which the proteins and enzymes involved in transcription bind to initiate the process is called a promoter. In most cases, promoters exist upstream of the genes they regulate. The specific sequence of a promoter is very important because it determines whether the corresponding gene is transcribed all of the time, some of the time, or hardly at all (Figure 2).
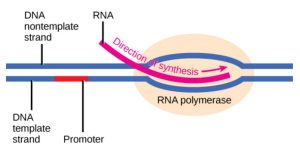
Figure 3.4: The initiation of transcription begins when DNA is unwound, forming a transcription bubble. Enzymes and other proteins involved in transcription bind at the promoter. OpenStax Concepts of Biology
Elongation
Transcription always proceeds from one of the two DNA strands, which is called the template strand. The mRNA product is complementary to the template strand and is almost identical to the other DNA strand, called the nontemplate strand, with the exception that RNA contains a uracil (U) in place of the thymine (T) found in DNA. During elongation, an enzyme called RNA polymerase proceeds along the DNA template adding nucleotides by base pairing with the DNA template in a manner similar to DNA replication, with the difference that an RNA strand is being synthesized that does not remain bound to the DNA template. As elongation proceeds, the DNA is continuously unwound ahead of the core enzyme and rewound behind it (Figure 3).
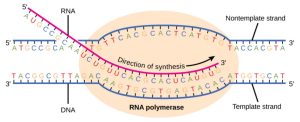
Figure 3.5: During elongation, RNA polymerase tracks along the DNA template, synthesizes mRNA in the 5′ to 3′ direction, and unwinds then rewinds the DNA as it is read. OpenStax Concepts of Biology
Termination
Once a gene is transcribed, the prokaryotic polymerase needs to be instructed to dissociate from the DNA template and liberate the newly made mRNA. Depending on the gene being transcribed, there are two kinds of termination signals, but both involve repeated nucleotide sequences in the DNA template that result in RNA polymerase stalling, leaving the DNA template, and freeing the mRNA transcript.
On termination, the process of transcription is complete. In a prokaryotic cell, by the time termination occurs, the transcript would already have been used to partially synthesize numerous copies of the encoded protein because these processes can occur concurrently using multiple ribosomes (polyribosomes) (Figure 4). In contrast, the presence of a nucleus in eukaryotic cells precludes simultaneous transcription and translation.
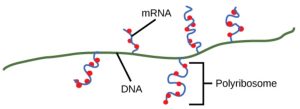
Figure 3.6: Multiple polymerases can transcribe a single bacterial gene while numerous ribosomes concurrently translate the mRNA transcripts into polypeptides. In this way, a specific protein can rapidly reach a high concentration in the bacterial cell. OpenStax Concepts of Biology
3.3 RNA Processing in Eukaryotes
The newly transcribed eukaryotic mRNAs must undergo several processing steps before they can be transferred from the nucleus to the cytoplasm and translated into a protein. The additional steps involved in eukaryotic mRNA maturation create a molecule that is much more stable than a prokaryotic mRNA. For example, eukaryotic mRNAs last for several hours, whereas the typical prokaryotic mRNA lasts no more than five seconds.
The mRNA transcript is first coated in RNA-stabilizing proteins to prevent it from degrading while it is processed and exported out of the nucleus. This occurs while the pre-mRNA still is being synthesized by adding a special nucleotide “cap” to the 5′ end of the growing transcript. In addition to preventing degradation, factors involved in protein synthesis recognize the cap to help initiate translation by ribosomes.
Once elongation is complete, an enzyme then adds a string of approximately 200 adenine residues to the 3′ end, called the poly-A tail. This modification further protects the pre-mRNA from degradation and signals to cellular factors that the transcript needs to be exported to the cytoplasm.
Eukaryotic genes are composed of protein-coding sequences called exons (ex-on signifies that they are expressed) and intervening sequences called introns (int-ron denotes their intervening role). Introns are removed from the pre-mRNA during processing. Intron sequences in mRNA do not encode functional proteins. It is essential that all of a pre-mRNA’s introns be completely and precisely removed before protein synthesis so that the exons join together to code for the correct amino acids. If the process errs by even a single nucleotide, the sequence of the rejoined exons would be shifted, and the resulting protein would be nonfunctional. The process of removing introns and reconnecting exons is called splicing (Figure 5). Introns are removed and degraded while the pre-mRNA is still in the nucleus.
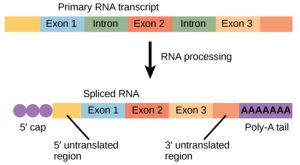
Figure 3.7: Eukaryotic mRNA contains introns that must be spliced out. A 5′ cap and 3′ tail are also added. OpenStax Concepts of Biology
3.4 Types of RNA involved in expression
In the complex world of molecular biology, RNA (ribonucleic acid) plays critical roles in gene expression, protein synthesis, and cellular regulation. There are various types of RNA that are involved in the process of gene expression, and they have different functions. In this chapter, we will explore the roles of mRNA, tRNA, rRNA, and snRNA, describing their structures, functions, and importance in the cell, and also discuss other types of RNA that are important for cell function and homeostasis.
Messenger RNA (mRNA)
Messenger RNA (mRNA) is a type of RNA molecule which is basically involved in protein synthesis as it works as a messenger between the DNA and the protein synthesizing apparatus of the cell. During transcription, mRNA is synthesized from a DNA template in the nucleus and then travels to the cytoplasm where proteins are synthesized. This molecule has three-base codons that pair with amino acids. The ribosomes, which are the cellular structures that are involved in the synthesis of protein, can translate the mRNA into a polypeptide chain of amino acids. In this way, mRNA acts as a translator as it connects the static genetic code present in the DNA to the dynamic machinery that is responsible for the construction of the proteins. This allows the genetic information that defines life to be translated into proteins that are vital in the growth, maintenance and function of living organisms.
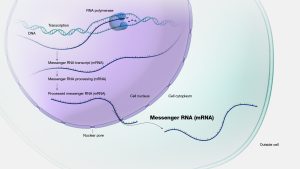
Figure 3.8: Structure and function of mRNA in protein synthesis. NHGRI
Ribosomal RNA (rRNA)
The molecular machines that make proteins, called ribosomes, depend on rRNA for both structural and functional support. rRNA and related ribosomal proteins are found in each of the two subunits that make up a ribosome. The ribosome is made up of two subunits: a 30S small subunit and a 50S large subunit in prokaryotes, and a 40S small subunit and a 60S large subunit in eukaryotes. For the ribosome to operate, rRNA molecules must fold into intricate three-dimensional structures.
In eukaryotes, there are four types of rRNA:
- 18S rRNA: A portion of the small 40S subunit.
- 28S rRNA, 5.8S rRNA, and 5S rRNA: A portion of the large 60S subunit.
In prokaryotes, there are three types of rRNA:
- 16S rRNA: A portion of the small 30S subunit.
- 23S rRNA and 5S rRNA: A portion of the large 50S subunit.
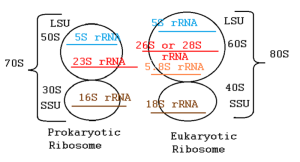
Figure 3.9: Comparison of prokaryotic and eukaryotic ribosomal subunits and associated rRNAs. Wikimedia
Function in Protein Synthesis: In the ribosome, rRNA has two primary functions:
- Structural support: It keeps the ribosomal subunits functional and intact.
2. Catalysis: During translation, rRNA catalyzes the formation of peptide bonds between amino acids. Since the polypeptide chain’s creation depends on rRNA’s catalytic activity, often referred to as peptidyl transferase activity, rRNA functions as a ribozyme, an RNA molecule with catalytic activity.
To ensure accurate translation of the genetic information, rRNA also makes sure that mRNA and tRNA align correctly during protein synthesis.
Transfer RNA (tRNA)
In the production of proteins, tRNA is essential. Amino acids are transported by tRNAs to the ribosome during translation, where they are incorporated into the expanding polypeptide chain. Every tRNA has an anticodon that connects with the matching codon on the mRNA strand and is unique to a specific amino acid. Intramolecular base pairing gives tRNA molecules their characteristic cloverleaf shape. This structure consists of:
- The amino acid attachment site at the 3′ end, where a specific amino acid is covalently attached by an enzyme called aminoacyl-tRNA synthetase.
- The anticodon loop, which contains a sequence of three nucleotides (the anticodon) that base pairs with the complementary codon on the mRNA during translation.
Since tRNAs are short, usually ranging from 76 to 90 nucleotides, their structure enables them to serve as adaptors between the relevant amino acids in the expanding polypeptide chain and the mRNA sequence. tRNAs move back and forth between the A (aminoacyl), P (peptidyl), and E (exit) sites of the ribosome during translation. A tRNA attaches to the mRNA codon in the A site. After the empty tRNA leaves through the E site, its amino acid is subsequently transferred to the expanding polypeptide chain in the P site. This procedure keeps going until a stop codon is reached, at which point a full protein is formed.
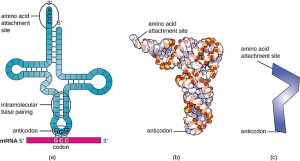
Figure 3.10: Structure and function of transfer RNA (tRNA). Wikimedia
Small Nuclear RNA (snRNA)
Small nuclear RNAs (snRNAs) are actively involved in the splicing of RNA which is the process of removing introns from pre-mRNA and joining the exons to form a mature mRNA transcript that can be translated into proteins. This process is crucial in order to produce functional mRNA that can then be used in protein synthesis. Being an essential part of spliceosome which is a large ribonucleoprotein structure that is present in the nucleus of eukaryotic cells, snRNAs control the splicing reactions. These RNAs are generally small and range from 100-300 nucleotides in length. They bind with certain proteins to form small nuclear ribonucleoproteins (snRNPs) which are the functional components of spliceosomes. The spliceosome relies on five key snRNAs for splicing: U1, U2, U4, U5, and U6. Each of these snRNAs contributes uniquely to the recognition of splice sites and the removal of introns.
snRNAs perform two primary roles during splicing by guiding the spliceosome to recognize conserved sequences at intron-exon boundaries:
- Intron Removal: snRNAs assist in correctly aligning the spliceosome at designated splice sites to excise introns.
- Exon Joining: snRNAs enable the ligation of exons, resulting in the formation of mature mRNA ready for nuclear export.
Mutations or defects in snRNAs or their associated proteins can lead to improper splicing, which is linked to various genetic diseases, such as spinal muscular atrophy.
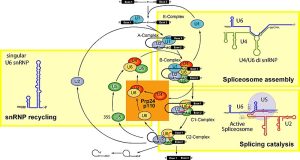
Figure 3.11: Spliceosome assembly and intron splicing mediated by snRNAs. Wikimedia
Other Important Types of RNA
MicroRNA (miRNA)
MicroRNAs (miRNAs) are small non-coding RNA molecules which are approximately 20-24 nucleotides in length that are involved in the regulation of gene expression at the post-transcriptional level. They base pair with target mRNAs which can lead to either degradation of the mRNA or translation inhibition. In this way, miRNAs allow for the adjustment of gene expression and thus contribute to cellular equilibrium. The roles of miRNAs are diverse in various biological processes, such as organismal development, cellular differentiation, apoptosis etc. The dysregulation of miRNA has been shown to be involved in several diseases, for instance, some miRNAs can act as oncogenes or tumor suppressors in cancer. Furthermore, miRNAs are also found to be promising biomarkers as well as therapeutic targets in disease management.
Small Interfering RNA (siRNA)
siRNAs are structurally related to miRNAs, and both are involved in the RNA interference (RNAi) process that is a gene-silencing mechanism. While miRNAs are known to have partial complementarity to their target mRNAs, resulting in gene silencing, siRNAs are designed to have complete complementarity to their target sequences which leads to mRNA degradation. This specificity renders siRNAs as a powerful research instrument and a potential candidate for therapeutic interventions especially in the modulation and suppression of genes implicated in various diseases. Scientists often utilize siRNAs in functional genomics studies to analyze gene roles by suppressing their expression. siRNAs are also being explored in clinical settings for the treatment of viral infections and cancer.
Long Non-Coding RNA (lncRNA)
lncRNAs are non-coding RNA molecules which are longer than 200 nucleotides and they do not code for proteins, but they are involved in the regulation of gene expression. These RNAs act as molecular scaffolds, guides, or decoys to interact with proteins and chromatin, influencing epigenetic modifications, transcriptional activity, and post-transcriptional processes. lncRNAs are pivotal in chromatin remodeling, transcriptional regulation, RNA splicing, and even nuclear architecture. They play a very important role in developmental biology and are being gradually acknowledged as fundamental components in diseases, including cancers and neurological disorders, as they can control tumor development or resistance to therapy.
Small Nucleolar RNA (snoRNA)
SnoRNAs are the small noncoding RNAs that are required for the chemical modification and processing of rRNAs and some other small RNAs. They guide specific modifications such as 2′-O-methylation and pseudo uridylation, which are critical for rRNA stability and ribosome functionality. Primarily located in the nucleolus, snoRNAs contribute to rRNA maturation and ensure the assembly of functional ribosomes. The alterations in snoRNA expression or dysfunction have been associated with genetic diseases such as dyskeratosis congenita and other ribosomopathies thus emphasizing the role of snoRNAs in cellular maintenance. New studies have also pointed towards the possibility of snoRNAs being involved in cancer and metabolic diseases.
- RNA Extraction and Purification
The accurate analysis of RNA starts at the proper extraction and purification. The isolation of RNA from microorganisms is a challenging process because of the presence of ribonucleases (RNases) that can lead to the degradation of RNA. Common methods for RNA extraction include:
Phenol-Chloroform Extraction: The method involves the use of acidic phenol and chloroform to achieve the separation of RNA from proteins and DNA. Trizol reagent is a well-known commercial material which is used for this purpose.
Column-Based Kits: These kits help in the process of RNA isolation where RNA is retained on a silica membrane while other components are washed off with salt and certain pH conditions. They provide enhanced reproducibility and are easy to use.
Magnetic Bead-Based Methods: Uses magnetic beads for the isolation of RNA and is especially helpful in high throughput screening.
Some of the difficulties encountered during the isolation of microbial RNA are the presence of thick cell walls in some microorganisms such as Gram-positive bacteria and fungi and this requires enzymatic or mechanical lysis for example using lysozyme or bead beating.
- Quantification and Quality Assessment
After extraction, RNA quality and quantity must be assessed to confirm that the sample is suitable for further processing:
Spectrophotometry:
- Nanodrop Spectrophotometer: Nanodrop measures RNA concentration based on absorbance at 260 nm. The 260/280 ratio assesses protein contamination, while the 260/230 ratio indicates organic solvent contamination.
- Qubit Fluorometer: Qubit uses dye-based fluorescence for precise RNA quantification, particularly for low-concentration samples.
Gel Electrophoresis:
- RNA integrity is measured by observing sharp 16S and 23S rRNA bands for bacteria or 18S and 28S rRNA bands for eukaryotic microbes. Smearing indicates degradation.
- RNA Sequencing (RNA-seq)
RNA sequencing has emerged as a powerful tool in the field of transcriptomics as it allows for the analysis of the entire set of RNA molecules in a microbial system. The workflow includes:
Library Preparation:
Conversion of RNA into cDNA through reverse transcription. Fragmentation and adapter ligation for making it compatible for sequencing.
High-Throughput Sequencing:
Some of the examples include the Illumina which produces millions of reads or the Oxford Nanopore.
Bioinformatics Analysis:
Mapping of the reads to the reference genomes or the de novo sequencing. Differential expression analysis is used to determine which genes are differentially expressed under various conditions.
Some of the applications of RNA-seq include analysis of microbial community shifts, stress response signaling and gene regulation networks.
- Northern Blotting
The northern blot, or RNA blot, is a technique used in molecular biology research to study gene expression by detection of RNA (or isolated mRNA) in a sample. Northern blotting technique still proves to be effective in identifying particular RNA molecules:
The RNA is first separated by gel electrophoresis and then transferred to a membrane.
The probes that are specific to the target RNA are hybridized and then detected which may involve the use of radioactive or chemiluminescent labels.
Although it is not as efficient as RNA-seq for getting the whole picture of the different transcripts, it is useful for the confirmation of specific transcripts.
Video:
- Reverse Transcription Quantitative PCR (RT-qPCR)
RT-qPCR is a sensitive method for quantifying RNA expression levels. The steps include:
- Reverse Transcription: In this process, RNA is first reversed into cDNA using reverse transcriptase.
- Quantitative PCR: This involves the amplification of the cDNA using certain primers and the quantification of the expression levels of the genes in real time.
Some applications are to confirm RNA-seq results and to assess the differential gene expression under various conditions such as exposure to certain environmental factors.
Videos:
- Emerging Techniques:
Single-Cell RNA Sequencing: It is a method of understanding RNA expression at the individual cell level which offers information on variation of cells present in microbial communities.
Video:
Direct RNA Sequencing: Comes with Oxford Nanopore and it is a method of sequencing RNA in its native state without the need for cDNA conversion and this means that modifications such as methylation are well preserved.
Video:
References:
https://commons.wikimedia.org/wiki/File:The_difference_between_ribose_and_deoxyribose.png
https://commons.wikimedia.org/wiki/File:RNA_structure_(full).png
https://openstax.org/books/concepts-biology/pages/9-3-transcription
https://commons.wikimedia.org/wiki/File:OSC_Microbio_11_04_tRNA.jpg
https://commons.wikimedia.org/wiki/File:Spliceosome_ball_cycle_new2.jpg
Alberts, B., Bray, D., Hopkin, K., Johnson, A.D., Lewis, J., Raff, M., Roberts, K., and Walter, P. (2015). Essential cell biology. Garland Science.
Bollu, A., Peters, A., and Rentmeister, A. (2022). Chemo-enzymatic modification of the 5′ cap to study mRNAs. Accounts of Chemical Research 55, 1249-1261.
Chen, Z.B., He, M., Li, J.Y.-S., Shyy, J.Y.-J., and Chien, S. (2023). Epitranscriptional Regulation: From the Perspectives of Cardiovascular Bioengineering. Annual Review of Biomedical Engineering 25, 157-184.
Greco, F.V., Pandi, A., Erb, T.J., Grierson, C.S., and Gorochowski, T.E. (2021). Harnessing the central dogma for stringent multi-level control of gene expression. Nature communications 12, 1738.

